| |
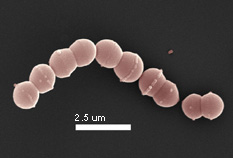 Photo: Robert
Hutkins, University of Nebraska
Photo: Robert
Hutkins, University of Nebraska |
Streptococcus thermophilus was
once described as a bacterium "marked more by the things which it
cannot do than by it’s positive actions" (Sherman, 1937). Although
it may be certainly be true that S. thermophilus is physiologically
and biochemically less versatile than other lactic acid bacteria, the reality
is that this organism can actually "do" quite a bit. In fact,
research during the past two decades has revealed that S. thermophilus has
properties that make it one of the most commercially important of all lactic
acid bacteria.
Streptococcus
thermophilus is used, along with Lactobacillus spp., as
a starter culture for the manufacture of several important fermented
dairy foods, including yogurt and Mozzarella cheese. Its use has
increased significantly during the past two decades, as a result
of the tremendous increase in consumption of these products. According
to USDA statistics, in 1998, more than 2.24 billion pounds Mozzarella
cheese and 1.37 billion pounds of yogurt were produced, respectively,
with a combined economic value of nearly $5 billion.
The substantial
increase in production of Mozzarella cheese and yogurt have led not
only to increased use of S. thermophilus cultures, but also
to new demands on their performance and production requirements. Industrial
strains, for example, should be insensitive to bacteriophage, have
stabile fermentation characteristics, and produce products having consistent
flavor and texture properties. Although research on the physiology
of S. thermophilus has revealed important information on some
of these properties, including sugar and protein metabolism, polysaccharide
production, and flavor generation, only recently has the genetic basis
for many of these traits been determined. Clearly, future efforts aimed
at improving this important industrial strain will require information
that can only be obtained by genome analysis.
Currently, several
traits in S. thermophilus have been targeted for strain improvement
programs (Delcour et al., 2000). Since bacteriophage are responsible
for considerable economic losses during cheese manufacture, efforts
are underway to engineer restriction and other phage resistance systems
into commercial strains. Enhancing stability and expression of exopolysaccharides
that act as natural thickening agents has also attracted significant
attention. Finally, S. thermophilus has an important role as
a probiotic, alleviating symptoms of lactose intolerance and other
gastrointestinal disorders.
The genome of S.
thermophilus is 1.8 Mb, making it among the smallest genomes
of all lactic acid bacteria. Although a moderate thermophile, it
is phylogenetically related to the more mesophilic lactococci and
has a comparable low G+C ratio (40%). Genes coding for metabolic
pathways involved in sugar catabolism (Poolman et al., 1989; Vaughan
et al., 2001), protein and peptide utilization (Fernandez-Espla et
al., 2000; Garault et al., 2002), polysaccharide production (Almirón-Roig
et al., 2000), the stress response system (Perrin et al., 1999),
and phage resistance mechanisms (Burrus, 2001; Solow and Somkuti,
2000) have been sequenced and characterized. More than 100 DNA sequence
entries are currently listed in GenBank. Although most strains do
not harbor plasmids, other mobile elements have been reported (Guedon
et. al., 1995), and techniques for gene transfer and mutagenesis
have been developed (Baccigalupi et al., 2000; Coderre and Somkuti,
1999). A genome sequencing project using an industrially-relevant
strain will undoubtedly reveal valuable information that could have
substantial impact on agriculture, the food industry, and the consuming
public.
References:
- Almirón-Roig,
E., F. Mulholland, M.J. Gasson, and A.M. Griffin. 2000. The complete
cps gene cluster from Streptococcus thermophilus NCFB 2393
involved in the biosynthesis of a new exopolysaccharide. Microbiol.
146:2793-2802.
- Baccigalupi,
L., G. Naclerio, M. de Felice, and E. Ricca. 2000. Efficient insertional
mutagenesis in Streptococcus thermophilus. Gene 258:9-14.
- Burrus, V.,
C. Bontemps, B. Decaris, and G. Guédon. 2001. Characterization
of a novel type II restriction-modification system, Sth368I, encoded
by the integrative element ICESt1 of Streptococcus thermophilus CNRZ368.
Appl. Environ. Microbiol. 67:1522-1528.
- Coderre, P.E.
and G.A. Somkuti. 1999. Cloning and expression of the pediocin operon
in Streptococcus thermophilus and other lactic fermentation
bacteria. Curr. Microbiol. 39:295-301.
- Delcour, J.,
T. Ferain, and P. Hols. 2000. Advances in the genetics of thermophilic
lactic acid bacteria. Curr. Opin. Biotechnol. 11:497-504.
- Fernandez-Espla,
M.D., P. Garault, V. Monnet, and E. Rul. 2000. Streptococcus thermophilus cell
wall-anchored proteinase: release, purification, and biochemical
and genetic characterization. Appl. Environ. Microbiol. 66:4772-4778.
- Garault, P.,
D. Le Bars, C. Besset, and V. Monnet. 2002. Three oligopeptide-binding
proteins are involved in the oligopeptide transport of Streptococcus
thermophilus. J. Biol. Chem. 277:32-39.
- Germond, J.E.,
M. Delley, N. D'Amico, and S.L. Vincent. 2001. Heterologous expression
and characterization of the exopolysaccharide from Streptococcus
thermophilus Sfi39. Eur. J. Biochem. 268:5149-5156.
- Guedon G., F.
Bourgoin, M. Pebay, Y. Roussel, C. Colmin, J.M. Simonet, and B. Decaris.
Characterization and distribution of two insertion sequences, IS1191
and iso-IS981, in Streptococcus thermophilus: does intergeneric
transfer of insertion sequences occur in lactic acid bacteria co-cultures?
Mol. Microbiol. 1995. 16:69-78.
- Perrin, C.,
C. Guimont, P. Bracquart, and J.L. Gaillard. 1999. Expression of
a new cold shock protein of 21.5 kDa and of the major cold shock
protein by Streptococcus thermophilus after cold shock. Curr.
Microbiol.39:342-347.
- Poolman, B.,
T. J. Royer, S.E. Mainzer, and B.F. Schmidt. 1989. Lactose transport
system of Streptococcus thermophilus: a hybrid protein with
homology to the melibiose carrier and enzyme III of phosphoenolpyruvate-dependent
phosphotransferase systems. J. Bacteriol. 171:244-253.
- Sherman, J.M.,
1937. The streptococci. Bacteriol. Rev. 1:3- 97. [NOTE: this was
the first review, in the first issue, of this seminal series of bacteriology]
- Solow, B.T.,
and G.A. Somkuti. 2000. Molecular properties of Streptococcus
thermophilus plasmid pER35 encoding a restriction modification
system. Curr. Microbiol. 42:122-128.
- Vaughan, E.E.,
P.T.C. van den Bogaard, P. Catzeddu, O.P. Kuipers, and W.M. de Vos.
2001. Activation of silent gal genes in the lac-gal regulon of Streptococcus
thermophilus. J. Bacteriol. 183:1184-1194.
|

