| |
 |
Cyanobacteria, because of their ability to photo-oxidize water, their nutritional simplicity, and
their amenability for genetic manipulation, offer great potential as cellular factories for biosolar
hydrogen production or alternatively for biomass-related energy applications. This proposal
seeks to determine the draft genome sequence of a biotechnologically important organism of
the phylum Cyanobacteria. This cyanobacterium is especially well suited to meet these goals.
Arthrospira maxima CS-328 (UTEX #2342) was isolated from Lake Chad in Chad, East Africa. It is
arguably the most robust oxygenic phototrophs known. Dismukes and coworkers have recently
shown (personal communication) that this cyanobacterium can produce hydrogen at rates similar
to those achieved with sulfur-starved Chlamydomonas reinhardtii. Arthrospira spp. are already
grown on an industrial scale, and an additional advantage of determining its genome sequence is
that this organism is already an important source of value-added biochemicals. This organism belongs to Section III of the
Cyanobacteria: filamentous organisms that do not produce heterocysts. At present, genomic
sequence information is available for only a single organism of this Section, Trichodesmium erythraeum. Thus, this sequence would significantly expand the genome database for this highly diverse group of phototrophs.
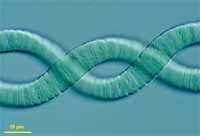
The genus Arthrospira comprises filamentous, non-heterocystous cyanobacteria that are
generally found in tropical and subtropical regions in warm bodies of water with high
carbonate/bicarbonate content, elevated pH, and salinity (3). Their large, gas-vacuolate filaments
(3 to 12 μm in diameter) are easily collected by filtration and other physical means of separation,
and blooms of these cyanobacteria have long been used as a food source by aboriginal peoples
around the world. For example, the Aztecs were exploiting this organism from Lake Texcoco in
Mexico before the arrival of Europeans (14). More recently, the phycologist Dangeard reported in
1940 that the Kanembu people in Chad consumed dihé, cakes of sun-dried cyanobacteria collected
from the shores of small ponds around Lake Chad (6, 7). While participating in a Belgian Trans-Saharan expedition 25 years later, the botanist Jean Leonard discovered a bloom of cyanobacteria
covering the waters around the shores of Lake Chad. He realized the connection between the
algal blooms and the dried dihé cakes sold in the markets. Arthrospira sp. additionally serve as the
principal food source for millions of filter-feeding flamingoes in the soda lakes of the Rift Valley
in central and eastern Africa. Carotenoids (principally echinenone, canthaxanthin, and
astaxanthin) derived from these cyanobacteria are responsible for the characteristic pink
coloration of the feathers of these birds, which are a principal source of food for migratory and
endemic raptors from Europe, Asia, and Africa. Arthrospira spp. are rich in amino acids, vitamins,
and γ-linolenic acid, and these bacteria are cultivated on an industrial scale as a source
4
of various specialty chemicals (e.g., β-carotene) and as a health food supplement under the name
“Spirulina.” 16S rRNA analyses indicate that the genera Arthrospira and Spirulina are not closely
related and that that the genus Arthrospira is a monophyletic group (2, 10).
References
1. Ananyev, G. and Dismukes, G. C. 2005. How fast can photosystem II split water? Kinetic
performance at high and low frequencies. Photosynth. Res. 84: 355-365.
2. Baurain, D., Renquin, L., Grubisic, S., Scheldeman, P., Belay, A., and Wilmotte, A. 2002.
Remarkable conservation of internally transcribed spacer sequences of Arthrospira
(“Spirulina”) (Cyanophyceae, Cyanobacteria) strains from four continents and of recent and 30-year-old dried samples from Africa. J. Phycol. 38: 384-393.
3. Boone, D. R. and Castenholz, R. W. (ed.) 2001. Bergey’s Manual of Systematic Bacteriology, 2nd Edition. Springer, Berlin.
4. Brown, I. I., Allen, C. C., Mummey, D. L., Sarkisova, S. A., and McKay, D. S. (2007). Irontolerant
cyanobacteria: implications for astrobiology. In: Extremophilic and Enigmatic Algae and
Non-Photosynthetic Protists, Seckbach, J. (ed.), Springer, Berlin.
5. Brown, I. I., Mummey, D. and Cooksey, K. E. 2005. A novel cyanobacterium exhibiting an
elevated tolerance for iron. FEMS Microbiol. Ecol. 52: 307-314.
6. Ciferri, O. 1983. Spirulina, the edible microorganism. Microbiol. Rev. 47: 551-578.
7. Ciferri, O and Tiboni, O. 1985. The biochemistry and industrial potential of Spirulina. Ann.
Rev. Microbiol. 39: 503-526.
8. Kawata, Y., Yano, S., Kojima, H., and Toyomizu. M. Transformation of Spirulina platensis
strain C1 (Arthrospira sp. PCC9438) with Tn5 transposase-transposon DNA-cation liposome
complex. Mar. Biotechnol. 6: 355-363.
9. Kruse, O., Rupprecht, J., Massgnug, J. H., Dismukes, C. G. and Hankamer, B. 2005.
Photosynthesis: a blueprint for solar energy capture and biohydrogen production
technologies. Photochem. Photobiol. Sci. 4: 957-969.
10. Manen, J.-F. and Falquet, J. 2002. The cpcB-cpcA locus as a tool for the genetic characterization
of the genus Arthrospira (Cyanobacteria): evidence for horizontal transfer. Int. J. Syst. Evol.
Microbiol. 52: 861-867.
11. Melis, A., Zhang, L., Forestier, M., Ghirardi, M. L., and Seibert, M. 2000. Sustained
photobiological hydrogen gas production upon reversible inactivation of oxygen evolution in
the green alga Chlamydomonas reinhardtii. Plant Physiol. 122: 127-136.
12. Pierson, B. K. and Parenteau, M. N. (2000) Phototrophs in high iron microbial mats:
microstructure of mats in iron-depositing hot springs. FEMS Microbiol. Ecol. 32: 181-196.
13. Pierson, B. K., Parenteau, M. N., and Griffin, B. M. 1999. Phototrophs in high-ironconcentration
microbial mats: physiological ecology of phototrophs in an iron-depositing hot
spring. Appl. Environ. Microbiol. 65: 5474-5483.
14. Sánchez, M., Bernal-Castillo, J., Rozo, C. and Rodríguez, I. 2003. Spirulina (Arthrospira): an
edible microorganism. A review. Universitas Scientiarum 8: 7-24. |

