Abstract
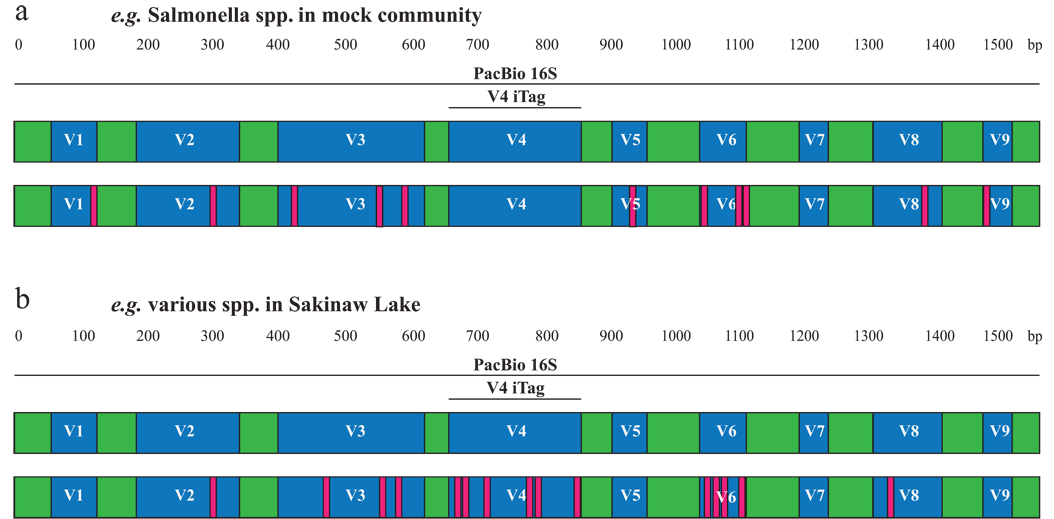
Over the past decade, high-throughput short-read 16S rRNA gene amplicon sequencing has eclipsed clone-dependent long-read Sanger sequencing for microbial community profiling. The transition to new technologies has provided more quantitative information at the expense of taxonomic resolution with implications for inferring metabolic traits in various ecosystems. We applied single-molecule real-time sequencing for microbial community profiling, generating full-length 16S rRNA gene sequences at high-throughput, which we propose to name PhyloTags.
We benchmarked and validated this approach using a defined microbial community. When further applied to samples from the water column of meromictic Sakinaw Lake, we show that while community structures at the phylum level are comparable between PhyloTags and iTags, variance increases with community complexity at greater water depths. PhyloTags moreover allowed higher unambiguous classification. Lastly, a platform-independent comparison of PhyloTags and in silico generated partial 16S rRNA gene sequences demonstrated significant differences in community structure and phylogenetic resolution across multiple taxonomic levels, including a severe underestimation in the abundance of specific microbial genera involved in nitrogen and methane cycling across the Lake’s water column. Thus, PhyloTags provide a reliable adjunct or alternative to cost-effective iTags, enabling more accurate phylogenetic resolution of microbial communities and predictions on their metabolic potential.