This genome was sequenced as a part of the large-scale multi-genome JGI CSP Saprotrophic Agaricomycotina Project (SAP), which focuses on the diversity and evolution of decay mechanisms, organismal phylogenetic relationships, and developmental evolution. A large collaborative effort led by PI of this project, David Hibbett (Clark University) aims for master publication(s) of the SAP data analysis. Researchers who wish to publish analyses using data from unpublished SAP genomes are respectfully required to contact the PI and JGI to avoid potential conflicts on data use and coordinate other publications with the SAP master paper(s).
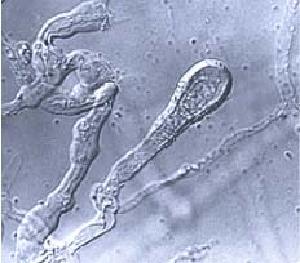
Phlebia brevispora Nakasone is a Basidiomycetes species that produces resupinate basidocarps on wood of conifers and hardwoods and is associated with a white rot. The species belongs in the phlebioid clade where many widespread white rot species are nested. Phanerochaete chrysosporium the model species for the study of white rot biochemistry is also nested in this clade. Different species of Phlebia have been demonstrated to produce lignin peroxidases, manganese peroxidases and laccases and glycoside hydrolases such as xylanases. The genome sequence of Phlebia brevispora will increase our knowledge on the white rot biochemistry of the phlebioid clade and it will be a great candidate for comparative studies with the genome sequence of Phanerochaete chrysosporium but also with the genome sequences of other white rot species. Furthermore the genome sequence will increase the opportunities for potential biotechnological applications of the species on bioremediation and bio-fuel technologies.
Genome Reference(s)
Binder M, Justo A, Riley R, Salamov A, Lopez-Giraldez F, Sjökvist E, Copeland A, Foster B, Sun H, Larsson E, Larsson KH, Townsend J, Grigoriev IV, Hibbett DS
Phylogenetic and phylogenomic overview of the Polyporales.
Mycologia. 2013 Nov-Dec;105(6):1350-73. doi: 10.3852/13-003
