Rhodopseudomonas palustris strains BisB5, HaA2, BisB18 and BisA53 (Bacteria, 5.5 Mb each): Phototrophic, fix carbon dioxide and nitrogen, produce hydrogen, degrade biomass-associated aromatic compounds under anaerobic (phototrophic) and aerobic (heterotrophic) conditions. One strain (CGA009) of R. palustris has been sequenced by the JGI and the sequence has revealed a surprising richness of metabolic versatility. This facultatively phototophic bacterium can grow with or without oxygen, fix nitrogen, produce hydrogen and it has highly developed biodegradation capabilities. The genome sequence of strain CGA009 revealed that the number of options that R. palustris has within its major metabolic modes to take advantage of fluctuating supplies of carbon, nitrogen, light and oxygen is even larger than was previously suspected based on physiological studies. It encodes for example, three nitrogenase isozymes, five benzene ring cleavage pathways and four light harvesting 2 systems. The genome sequence of strain CGA009 thus reinforces the notion that R. palustris is a generalist able to respond and adapt to changing conditions in the temperate soil and water environments in which it is so widely distributed. Against the background of general metabolic versatility of the R. palustris species there is considerable strain-to-strain diversity. Four additional strains of R. palustris will be sequenced. These additional genome sequences will provide a basis for defining the genetic and functional differences that can exist within the species Rhodopseudomonas palustris. The 16S rRNA sequence of each of the strains differs from that of the already sequenced strain (CGA009) by about 2%. The strains to be sequenced were isolated by direct plating from two different sites, one pristine and one polluted. The strains were frozen immediately following their isolation and have had minimal exposure to the laboratory environment. The strains to be sequenced differ from each other in BOX-PCR genomic DNA fingerprints. Although the strains have not been extensively characterized phenotypically, they have some obvious differences. They differ in pigmentation, in growth on benzoate, and the cells of each strain have slightly different shapes and sizes, although they all have the distinctive appearance of R. palustris. |
||
|
||
Rhodopseudomonas palustris BisA53

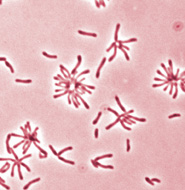 Rhodopseudomonas
palustris cells, credit. F. H. Harrison.
Rhodopseudomonas
palustris cells, credit. F. H. Harrison.