Silicibacter sp. TM1040 is a member of the Roseobacter clade of the alpha-proteobacteria, which is among the most abundant and ecologically relevant marine bacterial groups. Roseobacter isolates and gene sequences derived from them have been retrieved from marine environments ranging from sea ice to open ocean mixed layer to tropical coral reefs, and in ecological niches ranging from free-living plankton to sponge symbiont to biofilm pioneer. Although Roseobacter are cosmopolitan in the marine environment, their numbers and activity significantly rise with increases in the population density of phytoplankton and dinoflagellates. Some Roseobacter species have developed close associations with dinoflagellates and phytoplankton (1, 3) . Biological interactions between phytoplankton and bacteria may influence both the rate of primary production and the fate of fixed carbon in the surface ocean. Silicibacter sp. TM1040 was originally isolated as part of an investigation to understand the role of the bacteria in the physiology and toxigenesis of the dinoflagellate Pfiesteria piscicida (2) . Strain TM1040 is found within the phycosphere or physically attached to the surface of the dinoflagellate cell. Axenic cultures of P. piscicida grow poorly (and ultimately die) compared to cultures with associated bacterial assemblages, while adding back TM1040 restores normal growth (T.R. Miller, 2004, Ph.D. Dissertation). Although P. piscicida is a heterotrophic dinoflagellate, the Pfiesteria /TM1040 association is the only known "obligate" association between a dinoflagellate and a culturable bacterium. Additionally, TM1040 metabolizes the dinoflagellate secondary metabolite dimethylsulfoniopropionate (DMSP) via demethylation to methylmercaptopropionic acid (MMPA) (2) . DMSP is the major source of organic sulfur in the world's oceans and plays a significant role in the global sulfur cycle, facts that heighten the interest in Silicibacter sp. TM1040. TM1040 responds specifically to heat-labile compounds found in dinoflagellate homogenates, and is specifically chemotactic to DMSP and MMPA. The bacteria are also attracted to all amino acids, with the greatest response observed with methionine or valine (4) . These data suggest that TM1040 forms tight physical and physiological associations with its dinoflagellate host cell, in part through behavioral responses to dinoflagellate products, e.g., DMSP. An analysis of the TM1040 genome will potentially lead to better understanding of the cellular, physiological, and molecular mechanisms underlying this environmentally significant prokaryote and improve the understanding of the ecological interactions between bacteria and eukaryotic partner cells. This work also has the potential to significantly impact our understanding of the complex interactions between organisms important in influencing the sulfur and carbon cycles in the ocean. But the results from this research will also be of significant value to the scientific community, particularly current research efforts in microbiology, including those fields involved in signaling and sensory transduction, cell-cell interaction, and prokaryote:eukaryote interactions and communication. Thus, there is a potential to gain valuable new information about signaling between prokaryotes and eukaryotes through comparative analyses between the TM1040 genome and the genomes of other, free-living roseobacters. References 1. Alavi, M., T. Miller, K. Erlandson, R. Schneider, and R. Belas. 2001. Bacterial community associated with Pfiesteria -like dinoflagellate cultures. Environ. Microbiol. 3: 380-396. 2. Miller, T. R., and R. Belas. 2004. Dimethylsulfoniopropionate (DMSP) metabolism by Pfiesteria -associated Roseobacter spp. Appl. Environ. Microbiol. 70: 3383-3391. 3. Miller, T. R., and R. Belas. 2003. Pfiesteria piscicida, P. shumwayae , and other Pfiesteria -like dinoflagellates. Res. Microbiol. 154: 85-90. 4. Miller, T. R., K. Hnilicka, A. Dziedzic, P. Desplats, and R. Belas. 2004. Chemotaxis of Silicibacter sp. TM1040 towards dinoflagellate products. Appl. Environ. Microbiol. 70: 4692-4701.
|
||
|
||
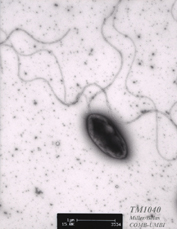
Silicibacter sp. strain TM1040