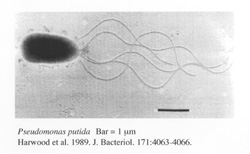
Pseudomonas putida F1 (Bacteria 6.2 Mb) is a versatile environmental isolate that is capable of growth on several aromatic hydrocarbons, including benzene, toluene, ethylbenzene and p -cymene. Its broad substrate toluene dioxygenase has been widely utilized in biocatalytic syntheses of chiral chemicals, as well as in the metabolism and detoxification of TCE, and in the production of indigo from indole. P. putida F1 is known to be chemotactic to aromatic hydrocarbons and chlorinated aliphatic compounds and has the potential for use in bioremediation applications. Members of the genus Pseudomonas are characterized by their ability to grow in simple media at the expense of a great variety of organic compounds. They have a strict respiratory metabolism and are motile by one or several polar flagella. Pseudomonas putida strains show optimal growth between 25-30?C and are easily isolated from soil and water by enrichment culture in mineral media with various carbon sources. Pseudomonas putida F1 was isolated from a polluted creek in Urbana, IL by enrichment culture with ethylbenzene as the sole source of carbon and energy. P. putida F1 is one of the most well-studied aromatic hydrocarbon degrading bacterial strains. Well over 200 articles have been written about various aspects of P. putida F1 physiology, enzymology, and genetics by microbiologists and biochemists, in addition to more applied studies by chemists and environmental engineers utilizing P. putida F1 and its enzymes for green chemistry applications and bioremediation. Strain F1 grows well with benzene, toluene, ethylbenzene, and p -cymene. Mutants of strain F1 that are capable of growth with n -propylbenzene, n -butylbenzene, isopropylbenzene and biphenyl are easily obtained. In addition to aromatic hydrocarbons, the broad substrate toluene dioxygenase in strain F1 can oxidize trichloroethylene (TCE), indole, nitrotoluenes, chlorobenzenes, chlorophenols and many other aromatic substrates. Although P. putida F1 cannot use TCE as a source of carbon and energy, it is capable of degrading and detoxifying TCE in the presence of an additional carbon source. The ability of P. putida F1 to degrade benzene, toluene, and ethylbenzene has a direct bearing on the development of strategies for dealing with environmental pollution. For example, many underground gasoline storage tanks are leaking and contaminating groundwater supplies with benzene, toluene, ethylbenzene and xylenes (BTEX). P. putida F1 can oxidize all of these compounds and thus a detailed knowledge of the physiology and biodegradation capabilities of this organism will essential for providing the scientific foundations for the emerging bioremediation industry.The sequence of F1 will provide an opportunity for comparative genomics of P. putida KT2440 and F1. As far as we know, P. putida KT2440 does not have the ability to grow on any aromatic hydrocarbons, although it has pathways for the degradation of numerous aromatic acids. Based on the significantly wider range of growth substrates known for P. putida F1, we expect to identify islands of catabolic diversity interspersed among standard P. putida genes. The KT2440 genome should provide a useful scaffold for the P. putida F1 genome sequence and this should speed the process of genome analysis and annotation. Regions of similarity should be immediately obvious, and annotation and functional genomic analysis can focus on unique regions of the sequence. Selected References 1. Choi, E. N., M. C. Cho, Y. Kim, K. C.-K., and K. Lee. 2003. Expansion of growth substrate range in Pseudomonas putida F1 by mutations in both cymR and todS , which recruit a ring-fission hydrolase CmtE and induce the tod catabolic pathway, respectively. Microbiology 149: 795-805. 2. Eaton, R. W. 1997. p -Cymene catabolic pathway in Pseudomonas putida F1: cloning and characterization of DNA encoding conversion of p -cymene to p -cumate. J. Bacteriol. 179: 3171-3180. 3. Gibson, D. T., J. R. Koch, and R. E. Kallio. 1968. Oxidative degradation of aromatic hydrocarbons by microorganisms I. Enzymatic formation of catechol from benzene. Biochemistry 7: 2653-2661. 4. Hudlicky, T., D. Gonzalez, and D. T. Gibson. 1999. Enzymatic dihydroxylation of aromatics in enantioselective synthesis: Expanding asymmetric methodology. Aldrichimica Acta 32: 35-62. 5. Lau, P. C. K., Y. Wang, A. Patel, D. Labbé, H. Bergeron, R. Brousseau, Y. Konishi, and M. Rawlings. 1997. A bacterial basic region leucine zipper histidine kinase regulating toluene degradation. Proc. Natl. Acad. Sci. USA 94: 1453-1458. 6. Parales, R. E., J. L. Ditty, and C. S. Harwood. 2000. Toluene-degrading bacteria are chemotactic to the environmental pollutants benzene, toluene, and trichoroethylene. Appl. Environ. Microbiol. 66: 4098-4104. 7. Wackett, L. P., and D. T. Gibson. 1988. Degradation of trichloroethylene by toluene dioxygenase in whole cell studies with Pseudomonas putida F1. Appl. Environ. Microbiol. 54: 1703-1708. 8. Zylstra, G. J., and D. T. Gibson. 1991. Aromatic hydrocarbon degradation: a molecular approach. Genet. Eng. 13: 183-203. 9. Zylstra, G. J., and D. T. Gibson. 1989. Toluene degradation by Pseudomonas putida F1: nucleotide sequence of the todC1C2BADE genes and their expression in E. coli. J. Biol. Chem. 264: 14940-14946. |
||
|
||
Pseudomonas putida F1