| |
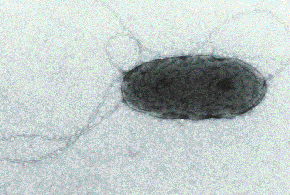
Photo: Yu.A.
Trotsenko and N.V. Doronina
|
The availability of
the genomic sequence for M. flagellatus, a b-proteobacterial methylotroph
will enhance and accelerate research in the fields of methylotrophy, microbial
diversity and metabolic evolution. Two methylotroph genome sequences are
currently available, covering two of the major classes of methylotrophs,
the a-proteobacterial serine cycle methylotroph Methylobacterium extorquens AM1
(Univ. of Washington; funded by NIH) and the the g-proteobacterial ribulose
monophosphate cycle methanotroph Methylococcus capsulatus Bath
(TIGR funded by DOE/Univ. Bergen). However, a significant gap in our genomic
level understanding of methylotrophy exists, as no sequence is available
for a representative of the third major group of methylotrophs, the ribulose
monophosphate pathway methanol-utilizers found in the b-proteobacteria.
This genome sequence will fill that gap and greatly enhance our ability
to analyze methylotrophy using genomic approaches. |