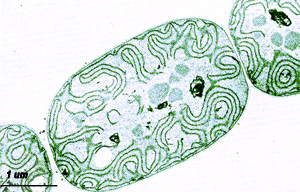
The primary reason to sequence the genome of this strain is to provide sequence data to relate to the many physiological, biochemical and genetic studies that have been and continue to be performed using this strain. In the 1970s, purified heterocysts from this strain were used to demonstrate that they are the sites of aerobic nitrogen fixation (Peterson and Wolk, 1978). More recently, Thiel (1999) has demonstrated multiple nitrogen fixation systems that are not present in any other commonly studied cyanobacterial strain. In addition, this strain produces H2 photosynthetically using solar power (Happe et al., 2000; Borodin et al., 2000). A. variabilis, like the other cyanobacterial strains sequenced by DOE, is important in global carbon sequestration via photosynthetic CO2 fixation. Because this strain can grow heterotrophically, it has been possible to create photosystem-I mutants of that grow in the dark (Nyhus et al., 1993). A. variabilis has additional energy-associated abilities. First, it is unique in having three different sets of nitrogenase genes that function under different environmental conditions. Two are molybdenum-dependent, one that functions exclusively in heterocysts (Thiel et al., 1995), whereas the second functions in vegetative cells, but only under anoxic conditions (Thiel et al., 1995, 1997a). Regulation of the latter nitrogenase genes is poorly understood. The third nitrogenase is vanadium-dependent but regulated by Mo (Thiel, 1993). When Mo is available, the heterocyst-specific Mo-nitrogenase(s) can function. When Mo is not available, the V-nitrogenase functions in heterocysts. The regulatory switch that controls V-nitrogenase expression is not understood. The genome sequence for A. variabilis will help us to take a systems biological approach to understanding the circuitry that controls the three nitrogenase systems and their relationship to Mo transport. The production of H2 by microorganisms has long been pursued actively. A practical process for biological H2 production using sunlight as the source of energy would create a major new source of sustainable and renewable energy, without greenhouse gas emissions or environmental pollution (Borodin et al., 2000). The laboratory of T. Happe studies the function of the uptake hydrogenase of A. variabilis in nitrogen and hydrogen metabolism using genetic techniques. A mutant in the hupSL operon encoding the uptake hydrogenase produces up to five times more hydrogen than the wild type (Happe et al., 2000). The genomic sequence of A. variabilis would aid in an understanding of the genes involved in H2 metabolism so that they could be manipulated for the bioengineering of strains that can produce large amounts of H2 photosynthetically. A third major area of interest in A. variabilis is comparative developmental analyses. Most cyanobacteria that form heterocysts also form spores, called akinetes (Adams and Duggan, 1999), that protect against desiccation, cold, and phosphate-deprivation. Essentially every vegetative cell of A. variabilis can differentiate into an akinete. Despite the structural and physiological differences of akinetes from heterocysts, their development sufficiently resembles the development of heterocysts that it has been proposed that heterocysts are evolutionary derivatives of akinetes (Wolk et al., 1994; Wolk, 1966). It is an ideal organism in which to pursue analysis of akinete formation, akinete-pattern formation, and the comparative differentiation processes of heterocysts and akinetes. The investigation of akinete formation in A. variabilis will profit greatly from the availability of genomic sequence data. References Adams DG, Duggan PS. (1999) Heterocyst and akinete differentiation in cyanobacteria. New Phytologist. 144: 3-33. Borodin VB, Tsygankov AA, Rao KK, Hall DO. (2000) Hydrogen production by Anabaena variabilis PK84 under simulated outdoor conditions. Biotechnology & Bioengineering 69(5, 5 Sep), 478-485. Happe T, Schütz K, Böhme H, (2000). Transcriptional and mutational analysis of the uptake hydrogenase of the filamentous cyanobacterium Anabaena variabilis ATCC 29413. J Bacteriol 182:1624-1631. Lang, N.J, Krupp, J.M., and Koller, A.L. 1987. J. Bacteriol. 169: 920-923. Electron micrograph of A. variabilis ATCC 29413. Nyhus KJ, Thiel T, Pakrasi HB. (1993) Targeted interruption of the psaA and psaB genes encoding the reaction-centre proteins of photosystem I in the filamentous cyanobacterium Anabaena variabilis ATCC 29413.Mol Microbiol. 9:979-88. Peterson, RB, Wolk, CP, (1978) High recovery of nitrogenase activity and of 55Fe-labeled nitrogenase in heterocysts isolated from Anabaena variabilis. Proc Natl Acad Sci USA. 75:6271-6275. Thiel T. (1993) Characterization of genes for an alternative nitrogenase in the cyanobacterium Anabaena variabilis. J. Bacteriology 175:6276-6286. Thiel T. (1999) Genetics and regulation of multiple nitrogenases in the cyanobacterium Anabaena variabilis ATCC 29413. pp. 1-27. In: Cyanobacterial and Algal Metabolism and Environmental Biotechnology. Edited by T. Fatma. Phycological Society. India. Thiel T, Lyons EM, Erker JC, Ernst A. (1995) A second nitrogenase in vegetative cells of a heterocyst-forming cyanobacterium. Proc. Natl. Acad. Sci. U.S.A. 92:9358-9362. Thiel T, Lyons EM, Erker JC. (1997) Characterization of genes for a second Mo-dependent nitrogenase in the cyanobacterium Anabaena variabilis. J. Bacteriol. 179:5222-5225. Wolk CP (1966) Evidence of a role of heterocysts in the sporulation of a blue-green alga. Am J Bot 53: 260-262. Wolk CP, Ernst A., Elhai J. (1994) Heterocyst metabolism and development, p. 769-823. In D. Bryant (ed.) Molecular genetics of cyanobacteria. Kluwer Acad Publ., Dordrecht, The Netherlands. |
||
|
||
Anabaena variabilis ATCC 29413