Within the framework of the JGI Mycorrhizal Genomics Initiative, we are sequencing a phylogenetically and ecologically diverse suite of mycorrhizal fungi (Basidiomycota and Ascomycota), which include the major clades of symbiotic species associating with trees and woody shrubs. Analyses of these genomes will provide insight into the diversity of mechanisms for the mycorrhizal symbiosis, including endo- and ectomycorrhiza.
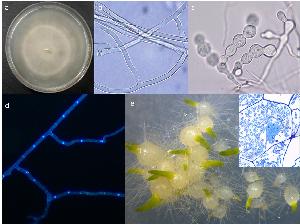
For their development, all orchids rely on the association with symbiotic fungi like Tulasnella calospora, that (at least in the early stages) provide the plant with organic carbon. T. calospora is the most common mycorrhizal partner of green orchids and belongs to the phylum Basidiomycota (order Cantharellales, family Tulasnellaceae). This fungus is distributed world-wide and normally found in every ecosystem, from tropical to temperate climate zones. Unlike some other symbiotic fungi identified in orchids, T. calospora can be easily cultivated in vitro. Thanks to these features T. calospora seems to be the best candidate model species for genome sequencing. In particular, T. calospora AL13/4D was isolated from mycorrhizal roots of the Mediterranean green orchid Anacamptis laxiflora. In our experiments, this strain proved to be the most effective in promoting symbiotic seed germination and plantlet development in vitro. Orchid symbiosis is still poorly studied, and T. calospora genome sequencing will allow us to make considerable advances in our understanding of the genetic and functional bases of the association, including carbon and nitrogen transfer to the host plant.
Genome Reference(s)
Kohler A, Kuo A, Nagy LG, Morin E, Barry KW, Buscot F, Canbäck B, Choi C, Cichocki N, Clum A, Colpaert J, Copeland A, Costa MD, Doré J, Floudas D, Gay G, Girlanda M, Henrissat B, Herrmann S, Hess J, Högberg N, Johansson T, Khouja HR, LaButti K, Lahrmann U, Levasseur A, Lindquist EA, Lipzen A, Marmeisse R, Martino E, Murat C, Ngan CY, Nehls U, Plett JM, Pringle A, Ohm RA, Perotto S, Peter M, Riley R, Rineau F, Ruytinx J, Salamov A, Shah F, Sun H, Tarkka M, Tritt A, Veneault-Fourrey C, Zuccaro A, Tunlid A, Grigoriev IV, Hibbett DS, Martin F
Convergent losses of decay mechanisms and rapid turnover of symbiosis genes in mycorrhizal mutualists.
Nat Genet. 2015 Apr;47(4):410-5. doi: 10.1038/ng.3223
