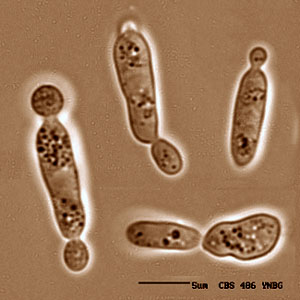
S.roseus is a unicellular basidiomycete "red" yeast species, a member of the class Urediniomycetes, that occurs in many different habitats but is frequently associated with plants. The attraction of S. roseus for genome sequencing is twofold. First, it has the smallest known genome size among basidiomycetes, by a factor of two. At 10 Mbp, the S. roseus genome is only 70% of that of the ascomycete Saccharomyces cerevisiae. The other Urediniomycete species whose genomes are currently being sequenced are plant pathogens (rusts and smuts) with genome sizes in the range of 25-90 Mbp and highly repetitive DNA content. Despite its small genome size, S. roseus is free-living and nonpathogenic. Sequencing the S. roseus genome will help identify the core gene set of basidiomycetes, assist with gene-finding in the plant pathogens, and contribute to our understanding of how genome size evolves. Second, the Urediniomycetes are among the most intron-riddled genomes known. Although only limited data is available for this group, it is not uncommon for a 3-Kbp gene to have 10 introns. The introns are typically about 80 bp long, and exons, 250 bp. Many of the introns in Urediniomycetes may have been recently inserted into their genes, which makes this group of fungi very interesting as a model for studying the evolutionary origins of introns. Sequencing the genome of S. roseus together with 3X coverage of Sporidiobolus salmonicolor should allow researchers to find new introns with identifiable sources.
