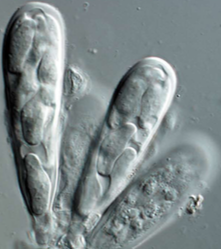
Image Credit: Pedro Crous
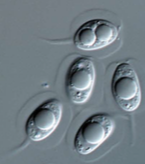
Image Credit: Pedro Crous
This genome was sequenced as part of the 1000 Fungal Genomes Project - Deep Sequencing of Ecologically-relevant Dikarya (CSP 1974) and more specifically as a part of the Dothideomycetes Sequencing Project, which seeks to densely sample members of a diverse lineage of saprotrophic, endophytic and pathogenic fungi to examine functional diversity of fungi with a shared evolutionary history.
Phyllosticta is an Ascomycete fungus in the Dothideomycetes clade. Phyllosticta capitalensis was described as a non-pathogenic endophyte with a wide host range and geographic distribution. Phyllosticta spp. have globally been recorded as endophytes, plant pathogens and saprobes from several plant hosts. Different Phyllosticta species have been associated with Citrus spp. in worldwide. Some of these cause important diseases such as Citrus black spot and Tan spot, subjected to phytosanitary legislation in the European Union and the U.S.A. Other species such as P. capitalensis are present only as endophyte on Citrus spp. Pathogenic species are often confused with a morphologically identical but non-pathogenic Phyllosticta species. Considering their economic importance, whole genome sequences for all the species involved with Citrus plants, including P. capitalensis, are needed to improve our understanding of the underlying differences in pathogenicity and their evolutionary separation. These data will also allow for the development of robust DNA barcodes for quick detection and will facilitate further research on this important Citrus pathogenic and non-pathogenic species.
Researchers who wish to publish analyses using data from unpublished CSP genomes are respectfully required to contact the PI and JGI to avoid potential conflicts on data use and coordinate other publications with the CSP master paper(s).
Genome Reference(s)
Guarnaccia V, Gehrmann T, Silva-Junior GJ, Fourie PH, Haridas S, Vu D, Spatafora J, Martin FM, Robert V, Grigoriev IV, Groenewald JZ, Crous PW
Phyllosticta citricarpa and sister species of global importance to Citrus.
Mol Plant Pathol. 2019 Dec;20(12):1619-1635. doi: 10.1111/mpp.12861
